
The primary emphasis of our research is building a successful breeding program though the development and application of translational genomic tools to improve selection efficiency for important traits in watermelon. Modern plant breeding has been revolutionized by the application of molecular tools to increase variation and selection efficiency. However, marker assisted selection (MAS) in watermelon has lagged behind, with most traits still being selected for based on the phenotype. This delays development of new cultivars with improved traits for growers and consumers.
Watermelon is an important crop worldwide and represents about 7% of world vegetable production acreage. In Georgia and the south eastern United States, the hot humid weather is conducive to the development of fungal diseases like Fusarium wilt and Gummy stem blight. Other traits of interest in watermelon breeding are earliness, fruit shape and size, sugar content and yield. In the United States, the crop is grown for its sweet edible flesh, but in other parts of the world the seed is also consumed as a source of oil and protein. In collaboration with industry and public partners we developed the first single nucleotide polymorphism (SNP) genetic maps for watermelon and identified the first quantitative trait loci (QTL) associated with many important traits. We are currently part of the CucCap project, a ~$7 million federally funded project to leverage genomics for improvement of disease resistance

Fusarium wilt is a major disease affecting watermelon production worldwide, including the southeastern United States. This soil borne disease is caused by Fusarium oxysporum f. sp. niveum (Fon) in watermelon. Management of the fusarium wilt is difficult because of the long-term survival of the pathogen in the soil, compounded by the phasing out of methyl-bromide leaving even fewer options for management of the disease. Our research focuses on identifying resistance to Fusarium wilt in watermelon crop wild relatives, and introgressing resistance loci into elite material.

This research was previously (2010-2015) funded through a USDA AFRI Research and Education Grant entitled “Translational Genomics For Enhancing Disease Resistance In Plants, An Internet-Facilitated Education Program For Training Plant Breeders” (GEO-2009-04819) and an USDA SCRI grant “A Systems Approach to Improve Disease Management and Production of Watermelon in the Southeastern U.S.” (2015-2018).
Current funding is provided by the UGA UGARF Cultivar development program.
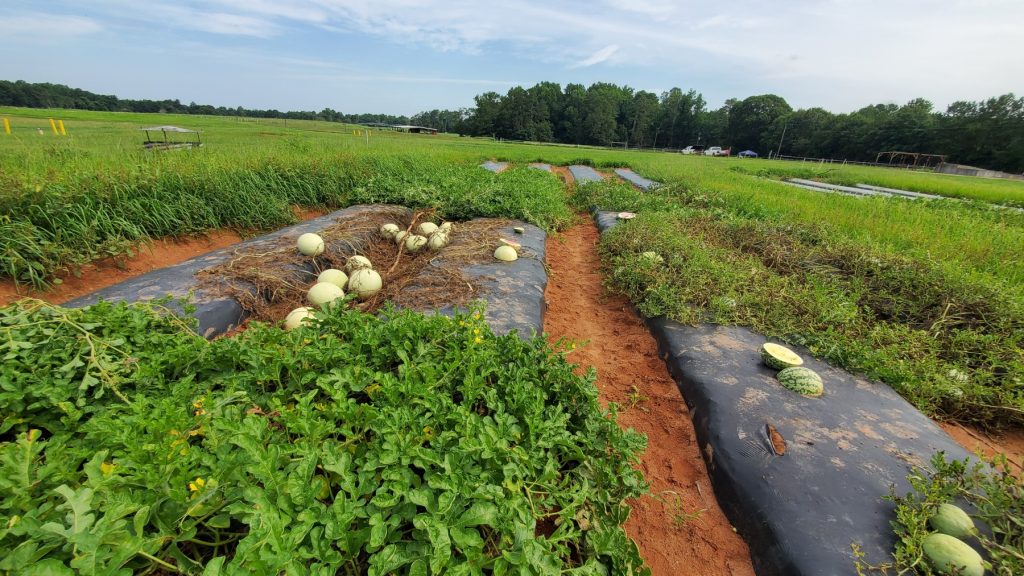



Gummy stem blight (GSB), is one of the most serious and devastating diseases of watermelon in the U.S. Even though resistant accessions have been identified many years ago, currently no resistant cultivars are available. Our lab is involved in mapping QTL assocuated with resistance to GSB and introgressing these loci into elite germplasm.
This research was previously (2010-2015) funded through a USDA AFRI Research and Education Grant entitled “Translational Genomics For Enhancing Disease Resistance In Plants, An Internet-Facilitated Education Program For Training Plant Breeders” (GEO-2009-04819).
Identification of QTL associated with gummy stem blight resistance in watermelon is part of the CucCap project funded by the USDA SCRI grant entitled “A Systems Approach to Improve Disease Management and Production of Watermelon in the Southeastern U.S.” (2015-2018)

In 2019 we launched a program to include breeding for whitefly (Bemisia tabaci) and whitefly transmitted viruses in Cucurbita pepo (squash and zucchini). In 2017 Cucurbit leaf crumple virus (CuLCrV; Begomovirus) and Cucurbit yellow stunting disorder virus (CYSDV: Crinivirus) were responsible for between 30% and 50% crop loss in squash and cucumbers in Georgia. No resistance for these viruses or whiteflies has been found in commercial cultivars tested to date. Currently the focus of our Cucurbita breeding program is identifying resistant germplasm. In collaboration with Dr. Geoffrey Meru at University of Florida, we are screening ~200 accessions from the USDA germplasm collection in the field in Tifton. The majority of these accessions are C. pepo, but other Cucurbita species are also included. A number of genotypes with resistance to CuLCrV and CuLCrV were identified in 2019 and are now being evaluated further for use in the breeding program. Once resistant germplasm is identified the resistance will be introgressed into elite squash and zucchini.
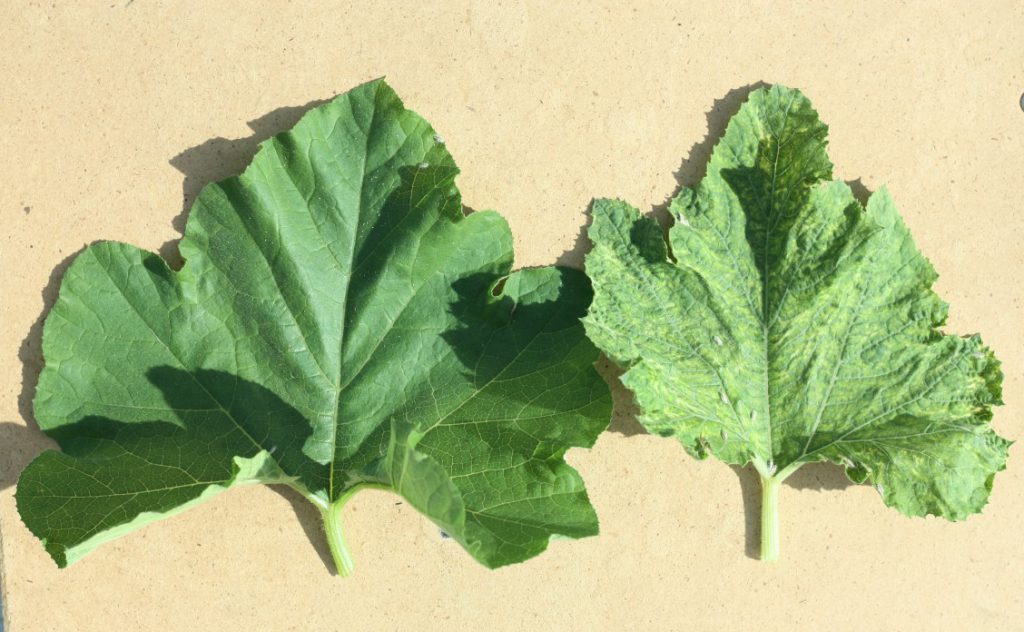



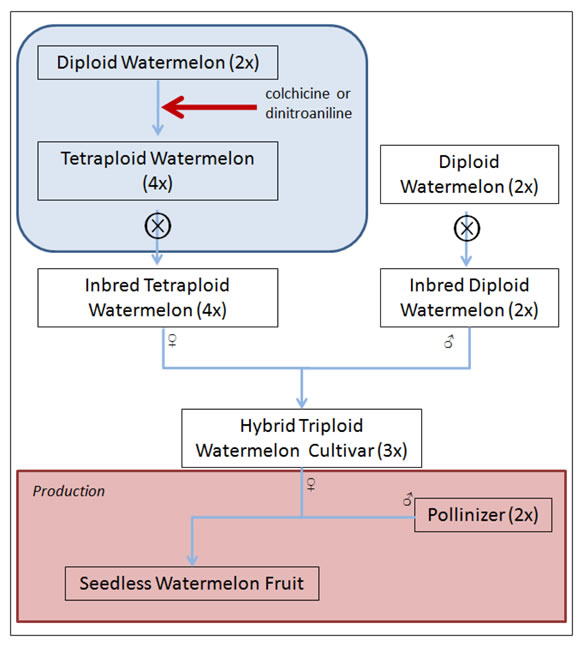
The percentage of watermelon production in the U.S. devoted to seedless fruit has increased dramatically from 51% in 2003 to 85% in 2009. Seedless fruit is produced on triploid plants, but because pollination is required for fruit set and triploid plants produce negligible amounts of viable pollen, diploid pollenizers are required. Due to preference for seedless fruit, U.S. watermelon production is dependent on synchronized flowering of diploid pollenizers and triploid watermelon cultivars for fruit production.
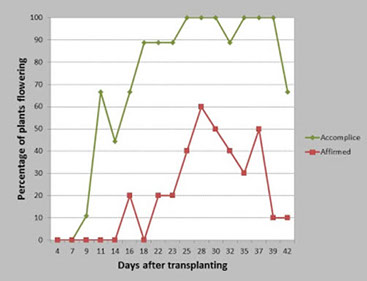
One of the key factors that determine flowering patterns, is time to first flower, measured in watermelon in days to male (DMF) and female (DFF) flowering. Very little was known about the control of flowering time in watermelon, with information lacking on the number of genes involved, mode of inheritance, heritability, and possible candidate genes. Our lab employed the translational genomic resources developed earlier by our group and determined that flowering time in watermelon is oligogenic with a major, stable, co-localized QTL on chromosome 3 responsible for ~50% of the phenotypic variation observed for DMF and DFF. This chromosomal region contains 172 genes, including homologues of the flowering locus T (Cla009504) and tempranillo 1 (Cla000855) genes associated with flowering time in other species. Currently we are investigating these candidate genes for functional markers for marker assisted selection of flowering time in watermelon.
We carried out a study to determine the overlap of male flowers produced by watermelon pollenizers with female flowers produced by triploid cultivars. You can use the Watermelon App to graph the flowering patterns for your chosen combinations.
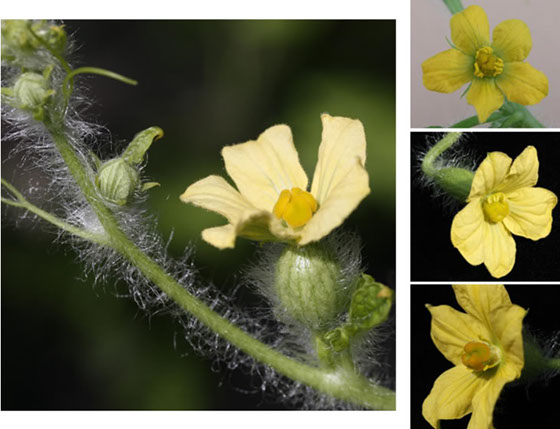
Sex expression is an important trait in watermelon where monoecious, andromonoecious and trimonoecious forms are present. The andromonoecious trait is highly undesirable in hybrid watermelon breeding programs since it requires emasculation during hybrid development. However this trait is desirable in pollenizer cultivars. In collaboration with Monsanto Vegetable Seeds, out lab identified four chromosomal regions associated with percent male (%M), percent female (%F), percent hermaphrodite (%HM) and percent female of pistillate (female + hermaphrodite; %F/P) flowers. The major QTL on chr. 3 is the expected location of the a gene. We determined that markers linked to two of the four QTL identified were located within 1 Mb of a 1-aminocyclopropane-1-carboxylic acid synthase (ACS) gene on the watermelon draft genome. ACS genes play a key role in sex expression in melon and cucumber, and these genes are therefore candidate genes for further studies to elucidate this trait in watermelon.

Consumer preference in the USA has shifted away from very large, picnic style watermelons to smaller blocky types. Watermelon size in the USA is classified as 60 count (9 ‐ 13.5 lbs), 45 count (13.6 ‐ 17.5 lbs), 36 count (17.6 ‐ 21.4 lbs) or 30 count (21.5+ lbs). We have identified several QTL associate with fruit size and shape, notably two major QTL on chromosomes 2 and 3. We have shown that the ClSUN25-26-27a (Cla011257), a member of the SUN gene family, is a candidate gene for the QTL on chromosome 3. Rind thickness is correlated with fruit size and consumers prefer thin rinds, but thicker rinds preferred for shipping. Currently we are identifying candidate genes for fruit size and shape and developing markers for marker assisted selection for these traits.
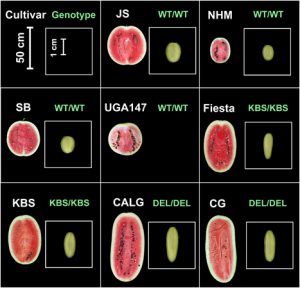
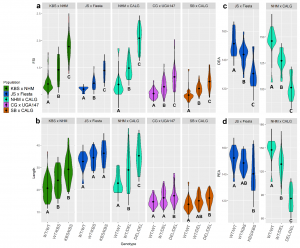
Seed size is important because watermelon breeders aim to develop hybrid cultivars with large seed for planting (especially where direct seeding is used), but that will produce fruit for eating with small seed. In the U.S., where seedless fruit are popular, this strategy is still followed in order to minimize the size of the white seed coat in the seedless fruit. In addition, in some parts of the world watermelon is grown for its edible seeds where a seed width of ~11 mm is desired for profitability.

Different species of edible seed watermelons (Citrullus spp.) are cultivated in Asia and Africa for their colorful nutritious seeds. Consumer preference vary for watermelon seed coat color. We mapped the QTL associated with seed coat color in watermelon.
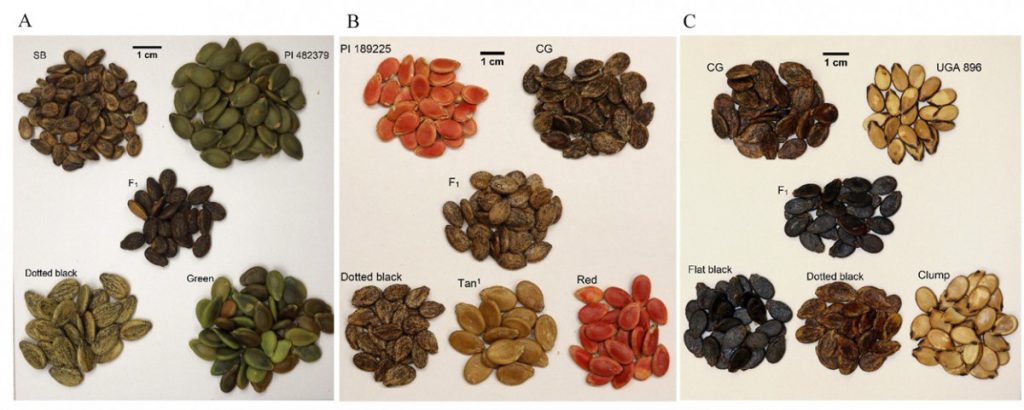

The egusi watermelon is popular in West Africa for its edible seed which is consumed in soups and stews. The seed can contain up to 40% seed oil, while the normal seed types contain ~25% seed oil. Genetic mapping studies have revealed that most of the variation in SOP between egusi and normal, non-egusi seed is explained by the egusi (eg) locus, which is also associated with the unique seed phenotype. Watermelon seed oil is high in unsaturated fatty acids, a profile similar to sunflower and soybean oil.

Diversification after domestication has led to cultivated watermelons exhibiting diverse fruit flesh colors, including red, yellow, and orange. Consumers prefer watermelon with sweet, red flesh, and the presence of lycopene, citrulline and arginine phytochemicals helpful for human health is an additional bonus. Breeders often select fruit with desirable flesh characteristics based on soluble solids content (Brix) and visual flesh color. We identified a region on chromosome 5 that is associated with sucrose, glucose, and fructose accumulation, while stable arginine content QTLs were identified on chromosomes 2 and 5.

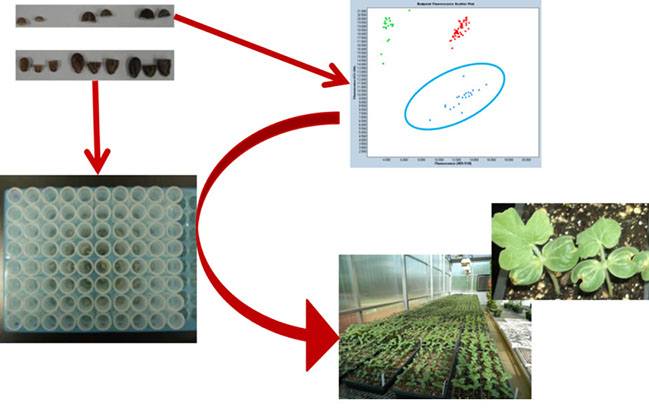
More than 90% of the watermelons grown in the United States are seedless cultivars. Seedless watermelons are produced through manipulation of ploidy and this breeding system poses challenges for marker assisted selection using current technology. Translating marker technology for use in triploid watermelon breeding is a focus of our current research.
Bell peppers have an annual farm gate value of over $100 million dollars in Georgia and are also some of the most popular vegetables grown by homeowners. The oomycete Phytophthora capsici causes root and fruit rot, stem blight and foliar blight in pepper causing global yield losses. Our lab used genotypic and phenotypic data to identify three accessions from resistant lines previously identified by Dr. Patrick Conner for use in breeding efforts to introgress resistance into bell peppers. In collaboration with Dr. Conner, we also identified genetic differences among seed sources of the heirloom pepper variety California Wonder. In collaboration with Dr. Pingsheng Ji and Dr. Suzanne O’Connell and a participatory plant breeding project at UGArden we launched a breeding effort for increased P. capsici resistance in bell peppers.


Funding



